Going through some old files I came across this piece of work we performed for a client some time ago -
Search Project
“…specific/sensitive biomarkers in certain cancers from the scientific literature and other online sources. For example might we be able to search in pancreatic or lung cancer for reports of biomarkers with say >70% sensitivity and/or specificity or maybe we could define an ROC of >0.75…
Of most interest would be a scan of the literature for all reported biomarkers in one or two cancers rather than just searching for CA19.9 data, for example. The objective for us would be to mine out the best biomarkers that we could then test and panel with our existing biomarkers.”
Robotic Reading
Here is what we were able to extract and auto-generate with a scan of the literature -
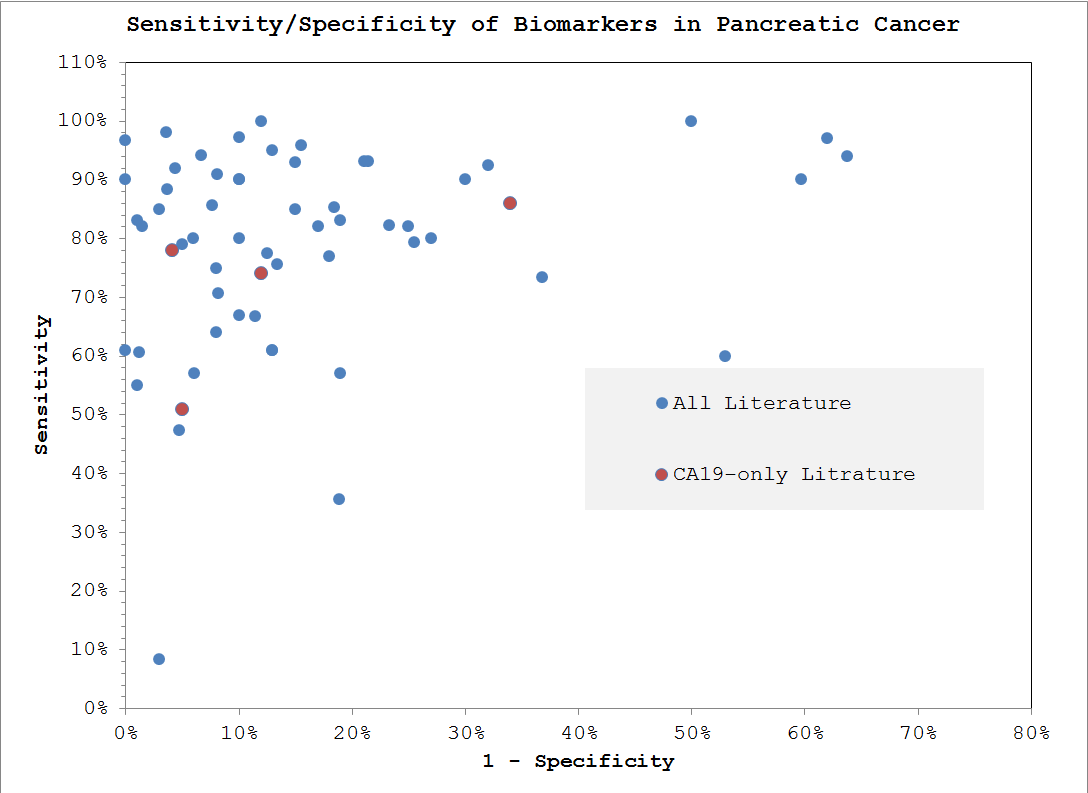
Design Choices
This wasn't a tricky project, so let us focus only briefly on some of the design choices.
Search Engine / Data Storage
We used an existing search engine to produce the corpus of material; this is very typical. The search engines do a great collation job, but we want to work with more than just a visual html response which is why we take this response several steps further.
In this case the data was sourced via PubMed, and we stored the search response in a simple database for us to add to, manipulate, and access as we wished at later stage.
Semantic Concepts
We created (and adjusted as more data arrived) certain semantic categories of idea, both by rule and by terminology. So for example if a term ended with a percentage sign that was treated as a 'SCORE'.
Similarly we created 'STATS' as a concept to catch any terminology using terms such as 'sensitivity', 'specificity' etc. Some other categories we created were DISEASE, CANCER, and SEARCH.
Categorisation primarily helps by broadening the nature of information extraction. That is, at some point you need to parse through all of the data 'noise' to extract your signal; categorisation helps us execute that parsing job in a complete - and organised - manner.
Regex for parsing
The response wasn't that large (perhaps 2000 articles) and, given the brief, we were able to use some fairly simple regular expressions in conjunction with our semantic categorisation to tease out the data we were looking for.
For the uninitiated, regular expressions, are a very fancy wildcarding mechanism. Regular Expressions can look really rather unsightly at first glance, a bit like hieroglyphics, but they get the job done.
If you are interested in learning more about regex check out one of the many cheatsheets, get golfing, or use this tool (one of my favourites) allowing you to experiment in real time.
Additional Analysis
In working with text one always tend to think of the body of material as one corpus and accordingly corpus analysis is something that happens very naturally.
In this instance we looked at neighbourhood/distance metrics for each document, i.e. what other papers read in a similar fashion to this particular one? To make that more concrete, here we can see the proximity of the first 5 neighbours of document 14:
In[8]: neighbours[14]
Out[8]:
[(0.17067703549375, Document(id='PFa8uOH-104', name=103)),
(0.15125379958999138, Document(id='PFa8uOH-418', name=417)),
(0.1370451828154102, Document(id='PFa8uOH-121', name=120)),
(0.13560045271802287, Document(id='PFa8uOH-130', name=129)),
(0.13023967761154998, Document(id='PFa8uOH-344', name=343))]
From here it is a short step to generating centrality measures and cluster metrics; and, although not part of this brief, to more advanced feature-extraction and classification.
... but, not Robotic Drawing
We can't quite claim that the project was entirely robotic-driven.
For the sake of completeness, I need confess that (for some unknown) we didn't fully automate the process: the bot produced the numeric results and dumped them into good 'ole fashioned excel which is where we created the graph.